Upperhost.com lists the best web hosting providers.  Michael L. Love Michael L. Love/proclus/GNU-Darwin link block
Related social networking sites that might be lesser known status and microblogs blogs bookmark sites Michael L. Love on Google Buzz Michael L. Love on Scientific Commons GNU-Darwin at eBay Just XI.NU!
Check out our Amazon Store,
or try the widget below.
Social Sharing |
Science Tools for GNU-DarwinDr. Michael L. Love,
Updates: |
  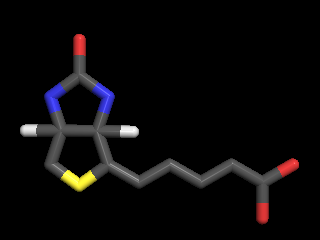 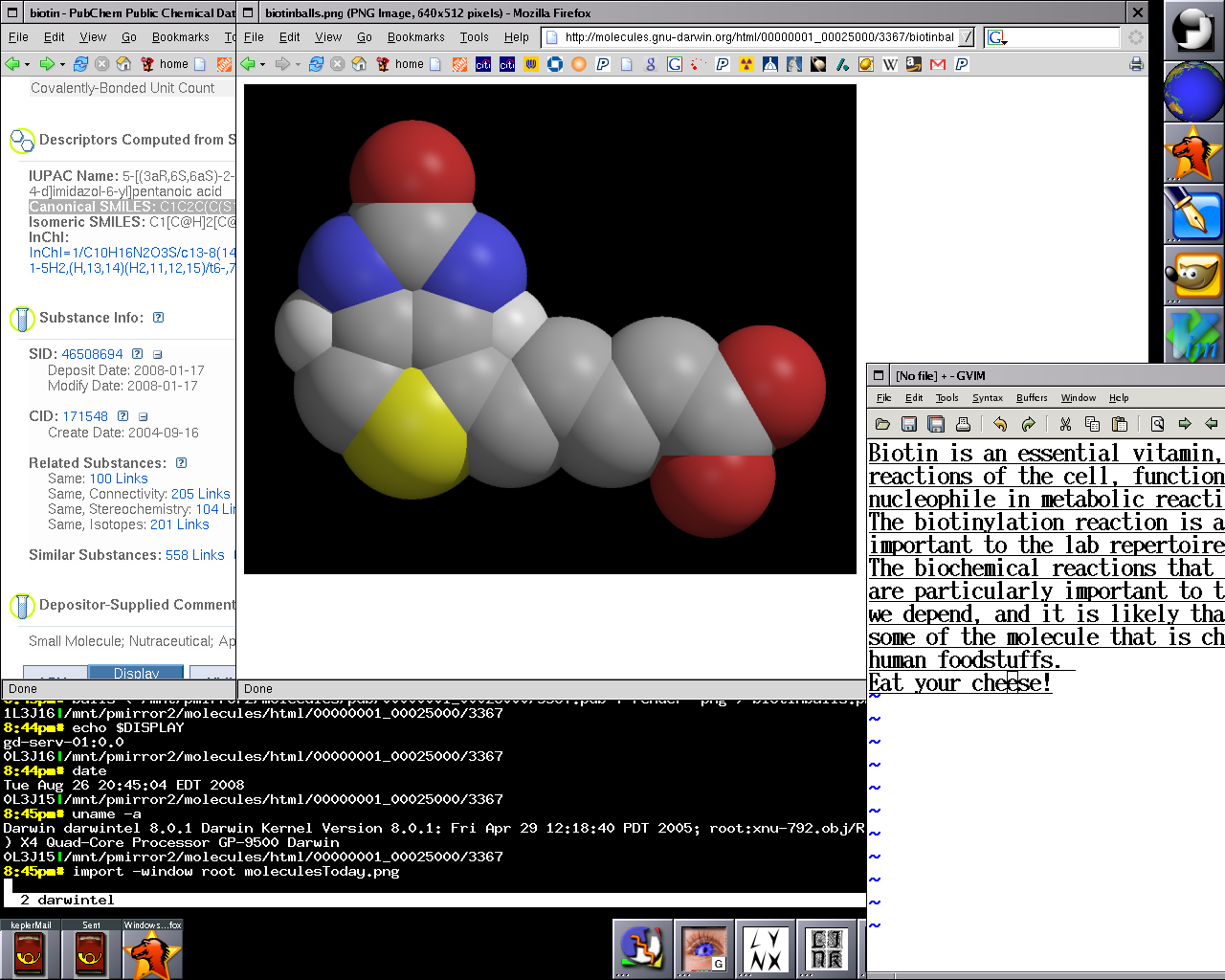 |
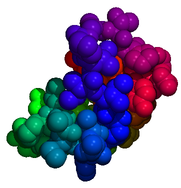
The Distribution is shepherding Molecules@GNU-Darwin.org,
a repository of over 4 million small molecule structural coordinate files in PDB format, which is
accessible by FOSS molecule viewer software that is provided by the Distro, such as PyMOL and RasMOL.
over 50 million structures are in the pipe. Molecules@GNU-Darwin.org demonstrates the kind of great
work that can be done in the scientific and education fields with GNU-Darwin OS and free and open source software.
There are also some notes about the package
set and Molecules machine. Molecules of the Day has an
RSS News Feed: |
GNU-Darwin hard media
|
Key scientific software packages are now available on our Office and Package discs.
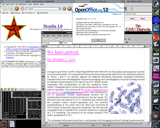
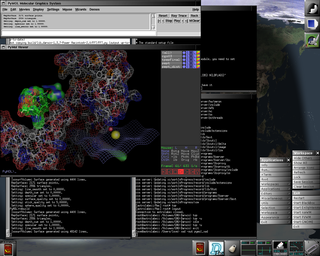
Favorite free software titles, first packaged for Darwin and Mac OS X by the Distribution, are included such as VIM, Ghostscript, Gnumeric, LaTeX, PyMOL (screenshot shown bottom left), Rasmol, gdFortran, LAM/MPI, AbiWord, GNUplot, and Raster3D. All are available for both x86 (Intel, AMD) and ppc (PowerPC) Darwin-only and Mac OS X systems.
Worry-free open source and command line installation is provided, and binary software distribution means that no compiling is necessary. Easy access to updates is also provided via our automated download system (ppc, x86).
As the premier free software distribution (founded Nov, 2000) for Darwin and Mac OS X, GNU-Darwin is clearly reliable with demonstrated stability and time-tested persistence. Our mature toolset has been successfully used for facilities administration, internet services, digital media production, intensive scientific calculation (see left center), authoring papers and progress reports (screenshot shown left top), high resolution graphics production, and all in a state of publication quality. With the assistance of a large open source development community, GNU-Darwin anticipates the needs of our users, and future Darwin utilities are provided in the present. GNU-Darwin is expressly engineered for high profile enterprises and cutting-edge scientific applications, but it is also suitable for workstation use and free software afficionados as well. Only with these discs, will users be up and running fast and getting real work done with all the desired tools, both now and in the future. |
GNU-Darwin hard media

Order them now!
GNU/Linux-x86 offerings
- A number of key software offerings have been prepared for use with the GNU/Linux User
Tarball. The package manager software for GNU/Linux is not yet ready for
production, but these files should be ready to use. Just untar them in
~/usr/local, and try them with the parallel
computing examples. PyMOL is
also included.
Please read the
alpha notice.
mpi-povray-3.1g.x86-linux.tar.gz
povray-3.1g.x86-linux.tar.gz
pvm-3.4.3.x86-linux.tar.gz
pvmpov-3.1e.2.x86-linux.tar.gz
pymol-0_78.x86-linux.tar.gz
If you have trouble with the User Tarball, try the latest files in CVS, or email proclus.
PyMOL for GNU-Darwin on Intel architecture (x86)
|
sample graphic installation script |
|
Announcing GNU-Darwin-OSF1 for Tru64 Alphas! This a0 distribution includes gdFortran, PyMOL, Python2.2, GCC, Raster3D, and many other great free software offerings. Please read the alpha notice.
Pymol has been built for Tru64 Alphas using the GNU-Darwin/OSF $HOME ports tarball. If you want to try it yourself, then read the GNU/Linux instructions and adapt them accordingly.
PyMOL-0.7.3 includes support for POV-ray, cartoon ribbons, transparency, and CCP4 maps. For the full update, including python-2.1.1, py-tkinter-2.1.1, povray-3.1g, and Mesa-4.0.1, run the following command.
- lynx -source
http://www.gnu-darwin.org/gdc/pymol073Update
| csh
|
|
|
Everyone please write to Stereographics development and sales to request Crystal Eyes for PyMOL and Apple computers. Read the letters from Drs. Love and Delano.
Donate: If GNU-Darwin is useful to you, then please consider contributing to the Distribution. Everyone can help.
Logan Donaldson has a new build of MOLMOL, which is based on Mesa-4.0.
We have created a number of Biology and Astronomy packages for Darwin-x86 users, gdFortran, Rasmol, and Raster3D included! More to come. Check out GNU-Darwin-x86, for Darwin/Intel.
Compiled Molscript sources are available from the Tenon third party apps page.
GNU-Darwin Fortran, has been updated for building CNS. Be sure to read the package installation instructions and check out Logan Donaldson's CNS notes. Read more.
Apple SciTech Group covered this development, which generated a little press.
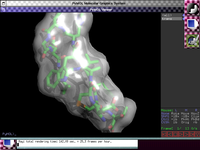
The PyMOL Molecular Graphics System has been ported to GNU-Darwin and Mac OSX.
|
PyMOL is a powerful molecular graphics package for structural biology
and other sciences. This interactive system is fully scriptable using the Python computing language,
and it is free software! PyMOL is in CVS,
and it will be added to the ports system.
Pymol is an effective molecular builder, but it can also produce stunningly beautiful and realistic 3D graphics. Be sure to see the GNU-Darwin Art page for an example. The GNU-Darwin build of PyMOL-0.60 is available now for distribution, along with the required dependent packages, tcl-8.3.2, tk-8.3.2, python-2.0.1, py-tkinter, and py-pmw. In addition, these packages also require XFree86-4.1 (see XonX). Be sure to read the package installation instructions. Incidentally, IDLE is a good editor/debugger for python scripts. IDLE is also available from GNU-Darwin. |
Other Key Crystallographic and Molecular Graphics Programs Ported *
CCP4-4.1 including PROTIN, Refmac, AMoRe, DM, MLPHARE, CAD, ARP_WARP, PLTDEV, FFT, OASIS, SCALEIT, SFALL, XDLMAPMAN, HKLVIEW, and RasMol
Here is a link to our 

 CCP4 porting tips page
CCP4 porting tips page

 .
.
CNS: no package available due to license restrictions.
- "YOU SHALL NOT DISCLOSE IN ANY FORM EITHER THE DELIVERED SOFTWARE OR
DOCUMENTATION OR ANY MODIFICATIONS OR DERIVATIVE WORKS BASED ON THE
SOFTWARE OR DOCUMENTATION TO THIRD PARTIES WITHOUT PRIOR WRITTEN
AUTHORIZATION."
POV-ray: with package available POV-ray builds automatically with the ports system.
- Be sure to consult the package installation
instructions.
PovScript: no package available due to license restrictions (see Molscript).
Molscript with GL: no package available due to license restrictions.
- "The LICENSEE shall not disclose in any form either the delivered
SOFTWARE or any modifications or derivative works based on the
SOFTWARE to third parties without prior written authorization from the
LICENSOR."
Tip: There are compiled molscript sources available from the Tenon third party apps page.
Raster3D: We now have a package available, and the Darwin modifications are in CVS
- Be sure to consult the package installation
instructions.
Mosflm: Here is a screen shot. Mosflm is now part of CCP4.
- Mosflm has been ported, but not yet tested to our knowledge.
DPS should be ready soon as well!
* The code will be optimized for the parallel architecture of an Apple G4 dual processor super cluster.
More Biology and Chemistry programs for GNU-Darwin
 William Van Etten of Blackstone Technology Group has kindly provided the
building instructions for EMBOSS, the sequence analysis
suite.
William Van Etten of Blackstone Technology Group has kindly provided the
building instructions for EMBOSS, the sequence analysis
suite.
babel: Molecular Coordinate Conversion Utility
clustalw: Multiple Sequence Alignment
fastdnaml: Maximum Likelihood Phylogenetic Tree Generation
gperiodic: GTK Periodic Table
hmmer: Markoff Analysis of Biological Sequences
molden: Molecular Graphics and Density Display
mopac: Calculation of Molecular Orbitals
nab: Nucleic Acid Building
ortep3: Thermal Ellipsoid Plotter for Crystal Structures
p5-bioperl: Perl Modules for Bioinformatics and Genomics
phylip: Package for the study of Phylogeny
povchem: PDB rendering for POVray
psi88: 3D Waveform Plotter
sim4: Alignment of Expressed and Genomic Sequences
xmolwt: Molecular Weight Calculator for Elemental Analysis
Astronomy Tools for GNU-Darwin
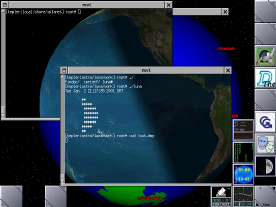
dgpsip: Internet DGPS corrections
luna: console moon phase calculator
p5-GPS: perl interface to a GPS receiver with Garmin protocol
rmap: Earth image generation
saoimage: Astonomical image display
sattrack: Satellite Tracking and Orbit Prediction Software
wmmoonclock: Moon Phase WindowMaker applet
wmspaceweather: WindowMaker space weather dockapp
wmsun: Sun rising and setting times WM dockapp
xearth: Earth imaging clock
xplanet: Planet rendering tool
Misc. Tools for GNU-Darwin
fftw: Discrete Fourier Transform calculation library
| Now you can do real science on your Mac! I have ported a variety of scientific computing programs to Darwin. In order to facilitate this work I have created a Package Distribution of Darwin and MacOSX software. |

















